3. General Run Parameters¶
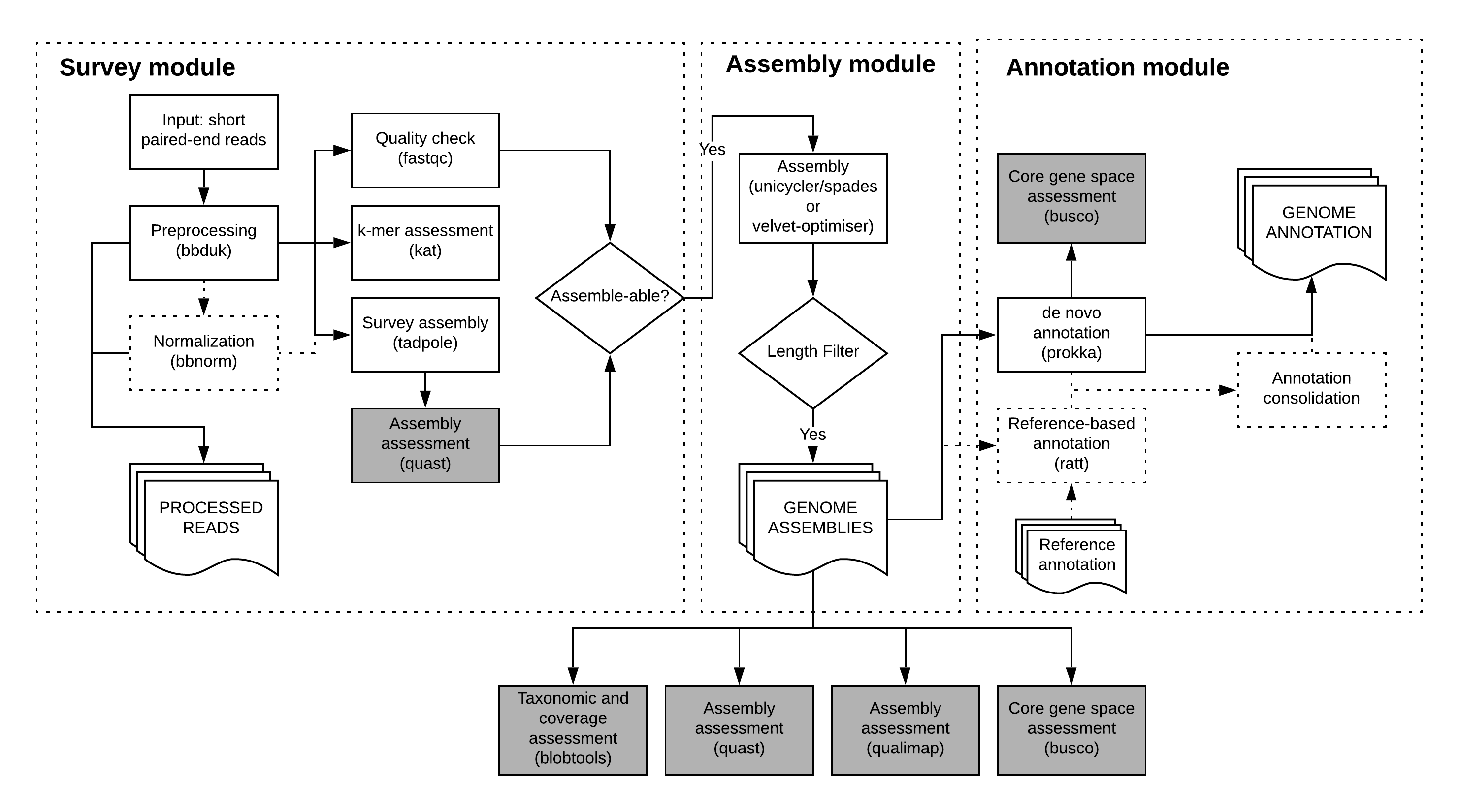
bgrr| consists of three main modules: survey, assemble, and annotate. Each module, or stage, has individual command line options. This section describes common command line options.
Each bgrr| run requires at the very least the following three command line parameters:
input_sheet--config--hpc_config
3.1. Command line arguments¶
The command bgrrl -h or bgrrl <stage> -h (or --help instead of -h) will display a list of command line options.
The first block of command line options is stage-specific and will be discussed in the description of each stage.
3.1.1. bgrr| options:¶
input_sheetThis is the path to a samplesheet, a comma-separated file, containing location and meta-information for each sample/library (REQUIRED!).
-o OUTPUT_DIR, --output-dir OUTPUT_DIROutput will be written to the specified directory [Analysis].
--project-prefix PROJECT_PREFIXReports and data packages will be prefixed by this string []
--config CONFIGPath to configuration file. This file specifies software configurations, resource locations and general project metadata (REQUIRED!).
--report-onlyWith this option, bgrr| Only (re-)runs reporting modules. No snakemake pipelines are called.
-f, --forceForce overwriting existing output directory, causes pipeline to be restarted. (CURRENTLY DISABLED)
--enterobase-groups ENTEROBASE_GROUPSComma-separated list of Enterobase microbial organisms. The set of assemblies is tested against organism-specific criteria and assemblies are packaged according to their species. [NEEDS REWORDING!]. By default, the enterobase mode is disabled.
3.1.2. HPC Options:¶
Controls for how jobs should behave across the HPC resources.
--partition PARTITIONWill run all child jobs on this partition/queue, this setting overrides anything specified in the
--hpc_configfile.--scheduler SCHEDULERThe job scheduler to use. LSF, PBS and SLURM are currently supported. If running without a scheduler type NONE here. Assumes SLURM by default.
--no_drmaaUse this flag if DRMAA is not available.
-N MAX_NODES, --max_nodes MAX_NODESMaximum number of nodes to use concurrently
-c MAX_CORES, --max_cores MAX_CORESMaximum number of cores to use concurrently
--hpc_config HPC_CONFIGConfiguration file for the HPC. Can be used to override what resources and partitions each job uses. (REQUIRED!)
--unlockIf the snakemake pipeline is not running because it is reporting that the directory is locked, then you can unlock it using this option. Please make sure that there are no other snakemake jobs are running in this directory before using this option!